- In-Stock Tumor Cell Lines
- Human Orbital Fibroblasts
- Human Microglia
- Human Pulmonary Alveolar Epithelial Cells
- Human Colonic Fibroblasts
- Human Type II Alveolar Epithelial Cells
- Human Valvular Interstitial Cells
- Human Thyroid Epithelial Cells
- C57BL/6 Mouse Dermal Fibroblasts
- Human Alveolar Macrophages
- Human Dermal Fibroblasts, Adult
- Human Lung Fibroblasts, Adult
- Human Retinal Muller Cells
- Human Articular Chondrocytes
- Human Retinal Pigment Epithelial Cells
- Human Pancreatic Islets of Langerhans Cells
- Human Kidney Podocyte Cells
- Human Renal Proximal Tubule Cells
What is MicroRNA
A microRNA is a small non-coding RNA molecule (containing about 22 nucleotides) found in plants, animals and some viruses. It functions in RNA silencing and post-transcriptional regulation of gene expression. miRNAs function via base-pairing with complementary sequences within mRNA molecules. As a result, these mRNA molecules are silenced, by one or more of the following processes:
- (1) Cleavage of the mRNA strand into two pieces,
- (2) Destabilization of the mRNA through shortening of its poly(A) tail,
- (3) Less efficient translation of the mRNA into proteins by ribosomes.
MiRNA Formation Process
A single pri-miRNA may contain from one to six miRNA precursors. These hairpin loop structures are composed of about 70 nucleotides each. Each hairpin is flanked by sequences necessary for efficient processing.
A nuclear protein recognizes the dsRNA structure of the hairpins in a pri-miRNA as DGCR8 or “Pasha”. DGCR8 associates with the enzyme Drosha, a protein that cuts RNA, to form the Microprocessor complex. In this complex, DGCR8 orients the catalytic RNase III domain of Drosha to liberate hairpins from pri-miRNAs by cleaving RNA about eleven nucleotides from the hairpin base (one helical dsRNA turns into the stem).
The product resulting has a two-nucleotide overhang at its 3′ end; it has 3′ hydroxyl and 5′ phosphate groups. It is often termed as a pre-miRNA (precursor-miRNA). Sequence motifs downstream of the pre-miRNA that are important for efficient processing have been identified.
In the cytoplasm, the pre-miRNA hairpin is cleaved by the RNase III enzyme Dicer. This endoribonuclease interacts with 5′ and 3′ ends of the hairpin and cuts away the loop joining the 3′ and 5′ arms, yielding an imperfect miRNA:miRNA* duplex about 22 nucleotides in length. Overall hairpin length and loop size influence the efficiency of Dicer processing.
The imperfect nature of the miRNA:miRNA* pairing also affects cleavage. Some of the G-rich pre-miRNAs can potentially adopt the G-quadruplex structure as an alternative to the canonical stem-loop structure. For example, human pre-miRNA 92b adopts a G-quadruplex structure which is resistant to the Dicer mediated cleavage in the cytoplasm.
Although either strand of the duplex may potentially act as a functional miRNA, only one strand is usually incorporated into the RNA-induced silencing complex (RISC) where the miRNA and its mRNA target interact.
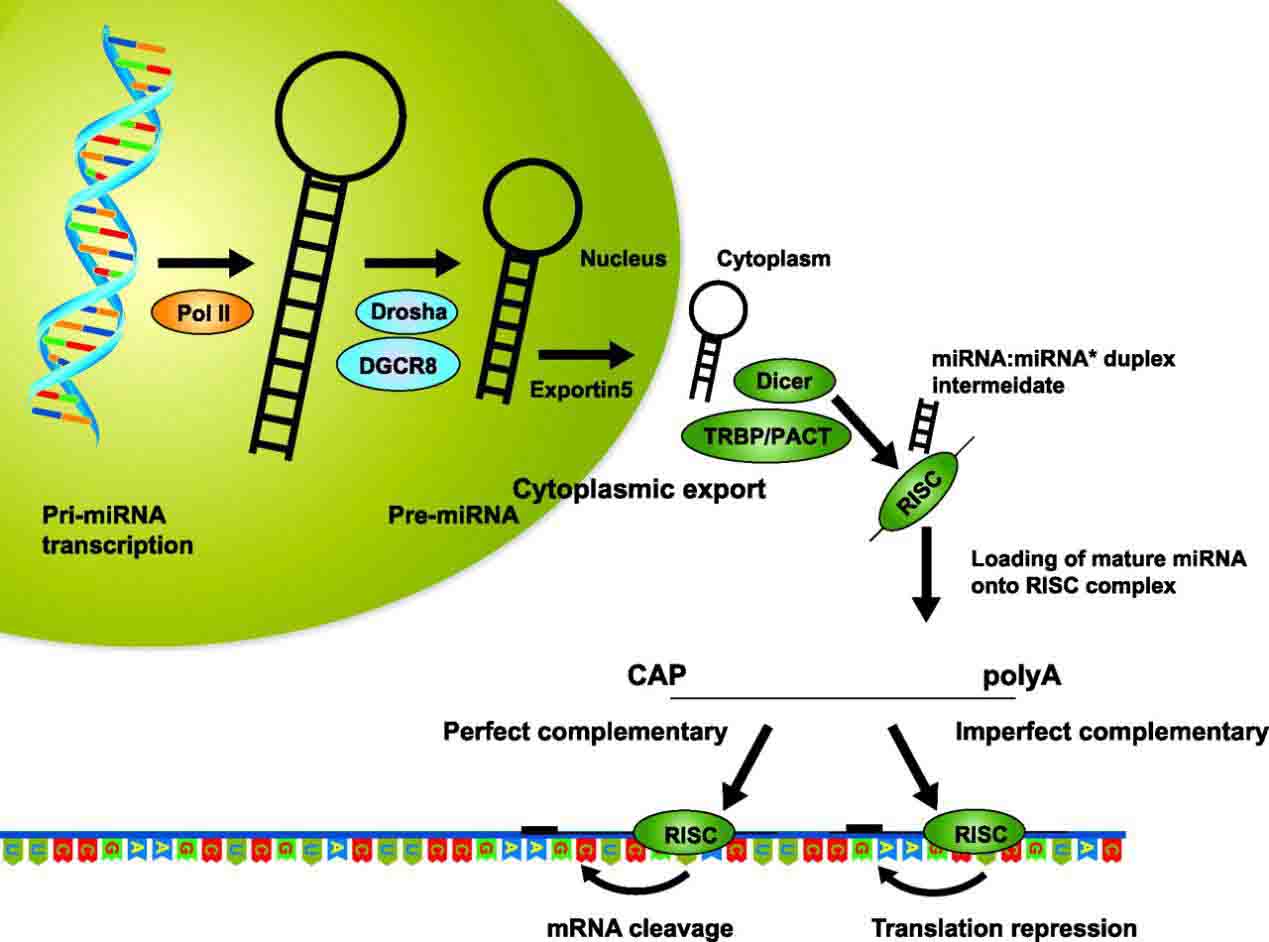
Fig.1 Nuclear and Cytoplasmic Processing
Nomenclature
Under a standard nomenclature system, names are assigned to experimentally confirmed miRNAs before publication. A dash and a number follow the prefix “miR”, the latter often indicating the order of naming.
For example, miR-124 was named and likely discovered before miR-456. A capitalized “miR-” refers to the mature form of the miRNA, while the uncapitalized “mir-” refers to the pre-miRNA and the pri-miRNA.
The miRNAs encoding genes are also named using the same three-letter prefix according to the conventions of the organism gene nomenclature.
For example, the official miRNAs gene names in some organisms are “mir-1 in C. elegans and Drosophila, Mir-1 in Rattus norvegicus and MIR-25 in humans.
MiRNAs with nearly identical sequences except for one or two nucleotides are annotated with an additional lower-case letter. Pre-miRNAs, pri-miRNAs, and genes that lead to 100% identical mature miRNAs but that are located at different places in the genome are indicated with an additional dash-number suffix.
Species of origin is designated with a three-letter prefix, e.g., hsa-miR-124 is a human (Homo sapiens) miRNA and oar-miR-124 is a sheep (Ovis aries) miRNA. Other common prefixes include ‘v’ for viral (miRNA encoded by a viral genome) and ‘d’ for Drosophila miRNA (a fruit fly commonly studied in genetic research).
MicroRNA Applications
It is only recently that researchers have begun to understand the scope and diversity of these regulatory molecules, even though the first miRNA was identified over ten years ago. More evidence shows that miRNAs exhibit a variety of crucial regulatory functions related to cell growth, development, and differentiation, and are associated with a wide variety of human diseases (cancer and heart disease).
Expression analysis studies reveal perturbed miRNA expression in tumors compared to healthy tissues. MicroRNAs are deregulated in breast, lung, and colon cancer, and upregulated in Burkitt’s and other human B-cell lymphomas.
As a consequence, human miRNAs are likely to be highly useful as biomarkers, especially for future cancer diagnostics, and are rapidly emerging as attractive targets for disease intervention.
In addition to their link with cancer, microRNAs play a significant role in the control of diverse aspects of cardiac function; and dysfunction as well. The misexpression of miRNAs is necessary and sufficient for multiple forms of heart disease, including myocyte growth, the integrity of the ventricular wall, contractility, gene expression, and maintenance of cardiac rhythm.
It is quite clear that miRNA research has great promise and may be the backbone of many of the most cutting-edge medical therapies of tomorrow. To keep up with the growing interest in miRNA, many miRNA products are emerging in the bio market, including miRNA isolation, amplification, profiling, and functional analysis; miRNA Agomir/Antagomir and services such as MicroRNA Agomir/ Antagomir Synthesis, libraries; An entire workflow of supporting reagents including cell culture, cell biology compounds and assays, transfection reagents, and much more.
AcceGen MicroRNA Products
AcceGen orders MicroRNA Agomir/Antagomir for gain-of-function studies or suppress microRNA activity. Use common miRBase identifiers to search for microRNAs in human (has-mir-), mouse (mmu-mir-), and rat (rno-mir-).
MicroRNA Agomir/ Antagomir Synthesis
AcceGen has pioneered the development of microRNA research and diagnostics tools with leading-edge services. We’re committed to pave new ways to meet all your needs for miRNA agomir/ antagomir synthesis.
AcceGen’s microRNA agomir/antagomir synthesis service has the following advantages:
- 1. Cover all human, mouse and rat miRNAs listed in miRbase
- 2. More stable, more manageable to pass the cell membrane and tissue gap
- 3. Purified and ready-to transfect
- 4. Short synthesis cycle and cost-efficiency
If you have any questions, please call us at 1-862-686-2696 or send an email to [email protected]. Your questions and requests will be answered by expert staff in AcceGen within 24 hours.

Copyright - Unless otherwise stated all contents of this website are AcceGen™ All Rights Reserved – Full details of the use of materials on this site please refer to AcceGen Editorial Policy – Guest Posts are welcome, by submitting a guest post to AcceGen you are agree to the AcceGen Guest Post Agreement – Any concerns please contact [email protected]