- In-Stock Tumor Cell Lines
- Human Orbital Fibroblasts
- Human Microglia
- Human Pulmonary Alveolar Epithelial Cells
- Human Colonic Fibroblasts
- Human Type II Alveolar Epithelial Cells
- Human Valvular Interstitial Cells
- Human Thyroid Epithelial Cells
- C57BL/6 Mouse Dermal Fibroblasts
- Human Alveolar Macrophages
- Human Dermal Fibroblasts, Adult
- Human Lung Fibroblasts, Adult
- Human Retinal Muller Cells
- Human Articular Chondrocytes
- Human Retinal Pigment Epithelial Cells
- Human Pancreatic Islets of Langerhans Cells
- Human Kidney Podocyte Cells
- Human Renal Proximal Tubule Cells
The Characteristics of COVID-19 Infection with Pneumonia
COVID-19 has greatly threatened human health worldwide due to the rapid spread of SARS-CoV-2, high morbidity, and mortality[1]. The clinical manifestations and spread of COVID-19 are complex[2]. Common symptoms include fever, cough, shortness of breath, chest pain, generalized fatigue, fatigue, headache, myalgia, insomnia, and diarrhea, while laboratory and radiological findings include lymphocytopenia and ground-glass opacities on chest imaging [3, 4].
COVID-19 pandemic is a global crisis that threatens our way of life[5]. As of June 2020, more than 350,000 people died from COVID-19. The global mortality rate of COVID-19 is about 7%, and the recovery rate is about 30%. Therefore, many doctors and scientists are working on coronavirus vaccines to overcome this disease.
Composition and Characteristics of Heparin
Heparin is the second most widely used drug in the world that has been safely used in medicine-cine for more than 80 years. It is formulated as a polydisperse heterogeneous natural product, including unfractionated heparin (UFH), low molecular weight heparin (LMWHs) and heparinoid. It is clinically approved as anticoagulants/thrombosis agent, with excellent safety, stability, bioavailability, and pharmacokinetic characteristics[6].
The Relationship between Heparin and COVID-19 Infected Pneumonia Patients
Patients with severe pneumonia caused by SARS-CoV2 have high platelet counts, and some patients have significant levels of D-dimer[7]. Thus, D-dimer may be an indicator for evaluating the prognosis of COVID-19 patients with high D-dimer using heparin. Heparin can prevent viral infections, including infections by members of the Coronavirus family[8]. As shown in Figures 1 & 2, Ning Tang et al. compared the 28-day mortality rate of patients who used heparin with those who did not use heparin in stratified patients. They found that patients who used heparin had a higher survival rate. Therefore, Patients with severe COVID-19 infection and D-dimer > 3.0 μg/mL have a higher survival rate and better prognosis after receiving heparin treatment [7, 9].

Figure1. Paired bar graphs showing mortality between heparin users and non-users in stratified
Patients [9].

Figure2. Paired bar chart showing the mortality between heparin users and nonusers in patients with severe pneumonia induced by SARS-CoV2. ( p<0.05)[7].
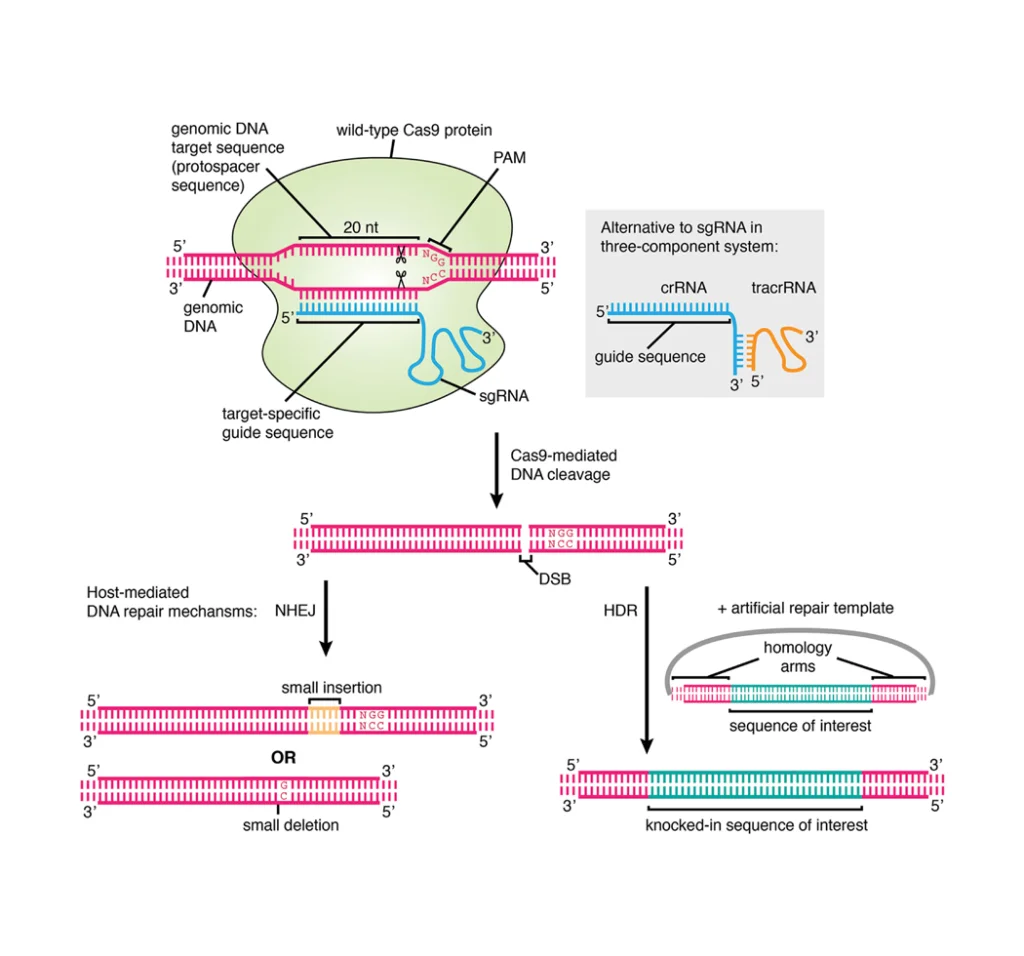
How to Prove that Heparin Inhibits the Virus from Entering the Cell?
According to reports, the spike protein of SARS-CoV-2 can bind to the receptor ACE2 on the surface of target cells. The spike protein has two binding domains: S1 has a high-affinity receptor-binding domain (RBD), and S2 has a host Sequence necessary for cell fusion. Heparin directly binds to S1 and interferes with the binding of SARS-CoV-2 to receptors on the surface of target cells to prevent new coronavirus from entering the cell [10].
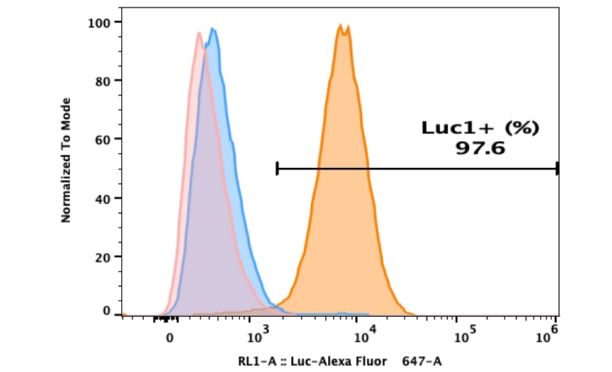
The researchers incubated the RT4 cell line (expresses both ACE2 and TMPRSS2) with heparin in combination with S1S2 at 37°C. As shown in Figure3, unfractionated heparin (UFH) at 10U/ml inhibited 80% of 330nM S1S2 binding to the cells and reached significance compared to untreated controls. [11].

Figure3. Unfractionated heparin inhibits S1S2 binding to RT4 cells(p<0.01)
RT4 cells were preincubated with the specified concentrations of unfractionated heparin, enoxaparin, and dalteparin at 37°C for 30 minutes, and then incubated with 100nM S1S2 at 37°C for another 60 minutes. After being placed at 37°C for another 60 minutes, the cells were washed and fluorescence-assisted anti-His6 was added for another 30 minutes at 21°C. Cell-related fluorescence was measured by flow cytometry and displayed as a percentage of S1S2 bound to untreated control cells. Their research shows that the two low molecular weight heparins (dalteparin and enoxaparin) are only partial inhibitors and not as effective as UFH[11].

Figure4. Concentration-dependent inhibition of S1S2 binding by unfractionated heparin and low molecular weight heparins, dalteparin, and enoxaparin.
Conclusion
It will cause a structural change in the spike protein when heparin binds to the spike protein of the SARS-CoV-2. Thus, affecting the binding of the virus to the receptor on the surface of the target cell and preventing the COVID-19 from entering the cell. At the same time, the heparin was used for patients with elevated D-dimer to increase their survival rate. Researchers can study the underlying mechanism of the existence of spike protein and use the biological indicators of elevated D-dimer to promote the development of related drugs to overcome COVID-19. Therefore, heparin may play an influential role in the treatment of patients with pneumonia caused by SARS-CoV-2 infection, which can reduce the mortality of patients.
AcceGen COVID-19 Research Related Cell Products
If your laboratory wants to achieve the same experiment, Human Bladder Cell Line – RT4 from AcceGen could be supportive.
Also, our featured cell lines: Huh7, Vero, Calu-3, HCT-8, and primary cells: Human Type II Alveolar Epithelial Cells, Human Pulmonary Alveolar Epithelial Cells, Human Bronchial Epithelial Cells, Human Nasal Epithelial Cells, have already applied to the research of coronavirus testing/infection and drug/vaccine development by researchers.
References
1. Muus C, Luecken MD, Eraslan G, Waghray A, Heimberg G, Sikkema L, Kobayashi Y, Vaishnav ED, Subramanian A, Smilie C et al: Integrated analyses of single-cell atlases reveal age, gender, and smoking status associations with cell type-specific expression of mediators of SARS-CoV-2 viral entry and highlights inflammatory programs in putative target cells. bioRxiv 2020.
2. Cao W, Li T: COVID-19: towards understanding of pathogenesis. Cell Res 2020, 30(5):367-369.
3. Xu Z, Shi L, Wang Y, Zhang J, Huang L, Zhang C, Liu S, Zhao P, Liu H, Zhu L et al: Pathological findings of COVID-19 associated with acute respiratory distress syndrome. The Lancet Respiratory Medicine 2020, 8(4):420-422.
4. Chaolin Huang* YW, Xingwang Li*, Lili Ren*, Jianping Zhao*, Yi Hu*, Li Zhang, Guohui Fan, Jiuyang Xu, Xiaoying Gu, Zhenshun Cheng, Ting Yu, Jiaan Xia, Yuan Wei, Wenjuan Wu, Xuelei Xie, Wen Yin, Hui Li, Min Liu, Yan Xiao, Hong Gao, Li Guo, Jungang Xie, Guangfa Wang, Rongmeng Jiang, Zhancheng Gao, Qi Jin, Jianwei Wang†, Bin Cao†: Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395(10223):497-506.
5. Alsamman AM, Zayed H: The transcriptomic profiling of COVID-19 compared to SARS, MERS, Ebola, and H1N1. bioRxiv 2020.
6. Mycroft-West CJ, Su D, Li Y, Guimond SE, Rudd TR, Elli S, Miller G, Nunes QM, Procter P, Bisio A et al: SARS-CoV-2 Spike S1 Receptor Binding Domain undergoes Conformational Change upon Interaction with Low Molecular Weight Heparins. bioRxiv 2020.
7. Yin S, Huang M, Li D, Tang N: Difference of coagulation features between severe pneumonia induced by SARS-CoV2 and non-SARS-CoV2. J Thromb Thrombolysis 2020.
8. Mycroft-West CJ, Su D, Li Y, Guimond SE, Rudd TR, Elli S, Miller G, Nunes QM, Procter P, Bisio A et al: Glycosaminoglycans induce conformational change in the SARSCoV-2 Spike S1 Receptor Binding Domain. bioRxiv 2020.
9. Tang N, Bai H, Chen X, Gong J, Li D, Sun Z: Anticoagulant treatment is associated with decreased mortality in severe coronavirus disease 2019 patients with coagulopathy. J Thromb Haemost 2020, 18(5):1094-1099.
10. Mycroft-West CJ SD, Pagani I, Rudd TR, Elli S, FGuimond SE: Heparin inhibits cellular invasion by SARS-CoV-2: structural dependence of the interaction of the surface protein (spike) S1 receptor binding domain with heparin. BioRxiv 2020.
11. Partridge LJ, Green LR, Monk PN: Unfractionated heparin potently inhibits the binding of SARS-CoV-2 spike protein to a human cell line. bioRxiv 2020.

Copyright - Unless otherwise stated all contents of this website are AcceGen™ All Rights Reserved – Full details of the use of materials on this site please refer to AcceGen Editorial Policy – Guest Posts are welcome, by submitting a guest post to AcceGen you are agree to the AcceGen Guest Post Agreement – Any concerns please contact marketing@accegen.com